Hsuan Megan Tsao, a PhD student in the Department of Human Genetics at McGill University received the Best PhD Oral Presentation Award at the Canadian Obesity Summit 2026, held in Montreal from March 25 to 29. The award recognizes outstanding research and excellence in scientific communication by an emerging leader in obesity science.
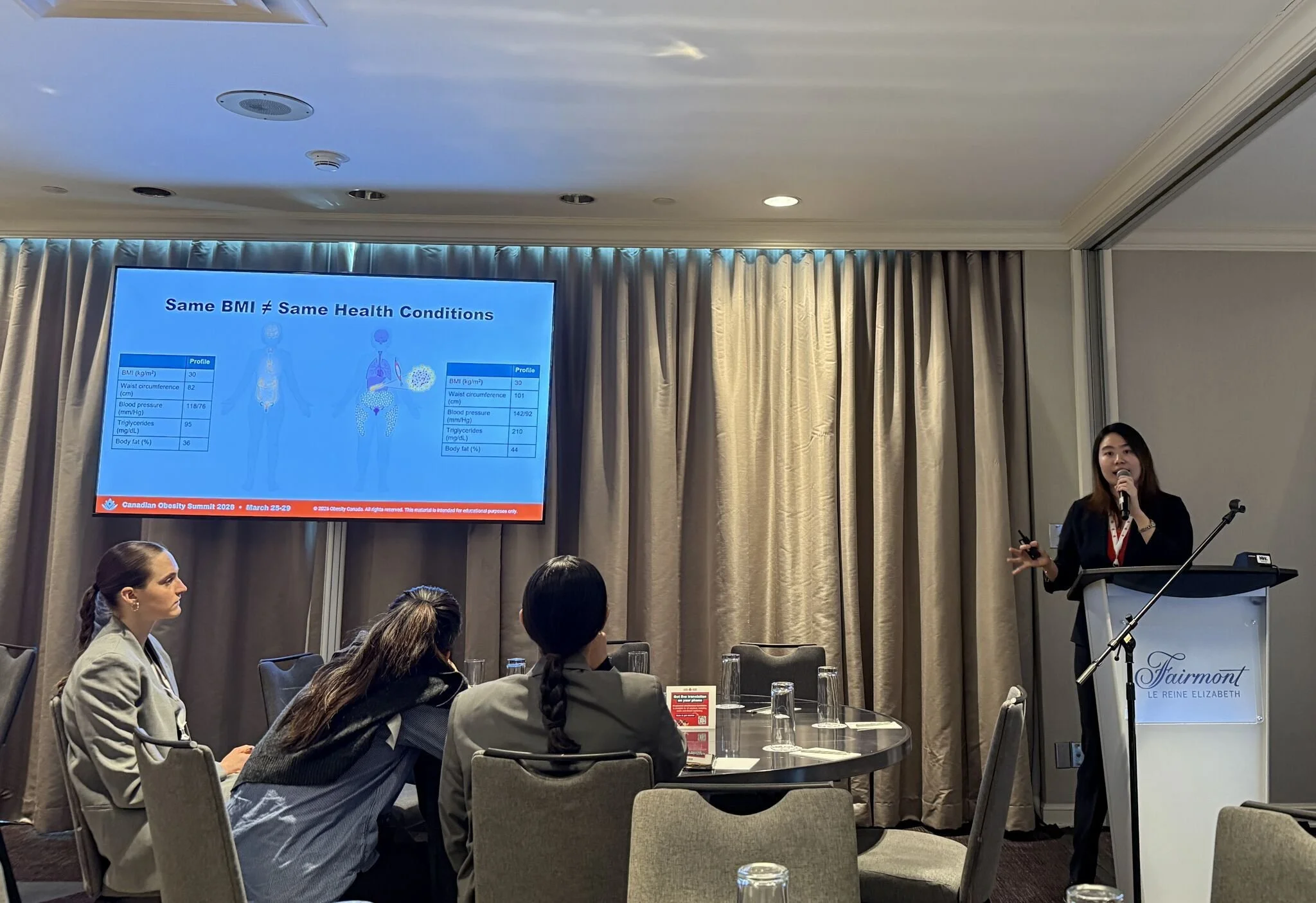
Tsao is supervised by Dr. Satoshi Yoshiji and is a member of the Canada Excellence Research Chair (CERC) in Genomic Medicine. She received the honour for her presentation, “Beyond BMI: Clinical Obesity Missed by BMI Carries a Threefold Higher Risk of Coronary Artery Disease.” Presented during the Summit’s scientific program, her work drew attention to a major limitation in current obesity assessment: the reliance on body mass index (BMI) alone to identify health risk.
Her research showed that some individuals who do not meet conventional BMI thresholds for obesity nonetheless carry substantially elevated cardiovascular risk. In particular, those with clinical obesity not captured by BMI had a threefold higher risk of coronary artery disease, highlighting how BMI-based approaches can miss patients who would benefit from earlier recognition and intervention.
The findings support a more clinically meaningful framework for obesity assessment that goes beyond weight alone to consider body composition, fat distribution, and obesity-related complications. Tsao’s presentation emphasized that individuals with similar BMI values can have markedly different cardiometabolic profiles, with important implications for screening, prevention, and treatment.
The judging committee recognized not only the significance of the research, but also Tsao’s ability to communicate complex clinical and genetic concepts with clarity, rigor, and practical relevance. The award was conferred by Canadian Obesity Summit leadership, including Dr. Sanjeev Sockalingam, Scientific Director; Nicole Pearce, Director of Education & Partnerships; and Lisa Schaffer, Executive Director.
Hosted by Obesity Canada, the Canadian Obesity Summit is the country’s leading conference dedicated to obesity research, education, and policy. The 2026 meeting brought together researchers, clinicians, trainees, and policymakers from across Canada to advance evidence-based obesity care.
Tsao’s recognition reflects the growing impact of McGill research and CERC in human genetics, cardiometabolic health, and precision medicine.
Advancing Precision Cardio‑Genomics: CERC Team Publishes New Findings in Circulation: Genomic and Precision Medicine











We are thrilled to highlight a new publication in Circulation: Genomic and Precision Medicine, authored entirely by members of our CERC community: Phenome-Wide Mendelian Randomization Identifying Circulating Proteins for Cardiovascular Traits in Populations of African Ancestry. This collaborative achievement underscores the program’s commitment to advancing equitable, genomics‑driven insights for global populations.
The article, “Phenome‑Wide Mendelian Randomization Identifying Circulating Proteins for Cardiovascular Traits in Populations of African Ancestry,” presents a rigorous and high‑resolution investigation of proteomic determinants of cardiometabolic disease, with a focus on populations historically underrepresented in genetic research.
About the Study
This study addresses a longstanding limitation in genomic medicine: the heavy bias toward discoveries in European‑ancestry cohorts. Using 2‑sample Mendelian randomization and colocalization analyses, the authors evaluated 1,562 circulating proteins across 145 cardiometabolic traits in individuals of African ancestry.
They then cross‑validated significant findings using large‑scale proteomic resources from the UK Biobank Pharma Proteomics Project, and further assessed whether signatures of natural selection may drive population‑specific protein–phenotype associations.
Key Discoveries
The team identified 115 robust protein–outcome associations in African‑ancestry populations, 51 of which differed significantly from European‑ancestry estimates—demonstrating ancestry‑specific biological effects that would have been overlooked in Euro‑centric datasets.
Four proteins—CD36, APOC1, GSTA1, and FOLH1—emerged as particularly compelling cardiometabolic targets, influencing lipid levels and heart‑related conditions in ways uniquely detectable in African‑ancestry cohorts.
The authors show that nearly 47.5% of these associations may be shaped by cis‑acting pQTLs under natural selection, highlighting evolutionary pressures as an underappreciated factor in precision medicine.
These findings not only broaden the catalog of actionable protein targets but also reveal how population‑specific variation can reshape the landscape of drug discovery, risk prediction, and mechanistic understanding of cardiometabolic disease.
Why This Matters for Precision Medicine
This research exemplifies why genomic diversity is essential for developing equitable therapies. Ancestry‑aware analyses can identify biomarkers and therapeutic targets that benefit patient groups routinely excluded from large‑scale studies—supporting the development of treatments that work for everyone, not just the populations most represented in research.
CERC Team Contribution
All authors of this publication are members of the Canada Excellence Research Chair in Genomic Medicine program. Their collective expertise—spanning proteomics, statistical genetics, cardiometabolic genomics, and evolutionary biology—made it possible to produce this ambitious and impactful analysis.
This work represents yet another step forward in CERC’s mission to integrate multi‑omics, population genetics, and precision health for global impact.
New Insights into GLP‑1RA Cardiometabolic Benefits Highlighted in Recent CERC‑Supported Publication
We are pleased to highlight an important new contribution from our CERC community: the publication of “Cardiovascular risk reduction with glucagon‑like peptide‑1 receptor agonists is proportional to HbA1c lowering in type 2 diabetes: An updated meta‑regression analysis incorporating FLOW and SOUL trials” in Diabetes, Obesity and Metabolism. This work features key contributions from Masashi Hasebe, Satoshi Yoshiji, and Chen‑Yang Su.
About the Study
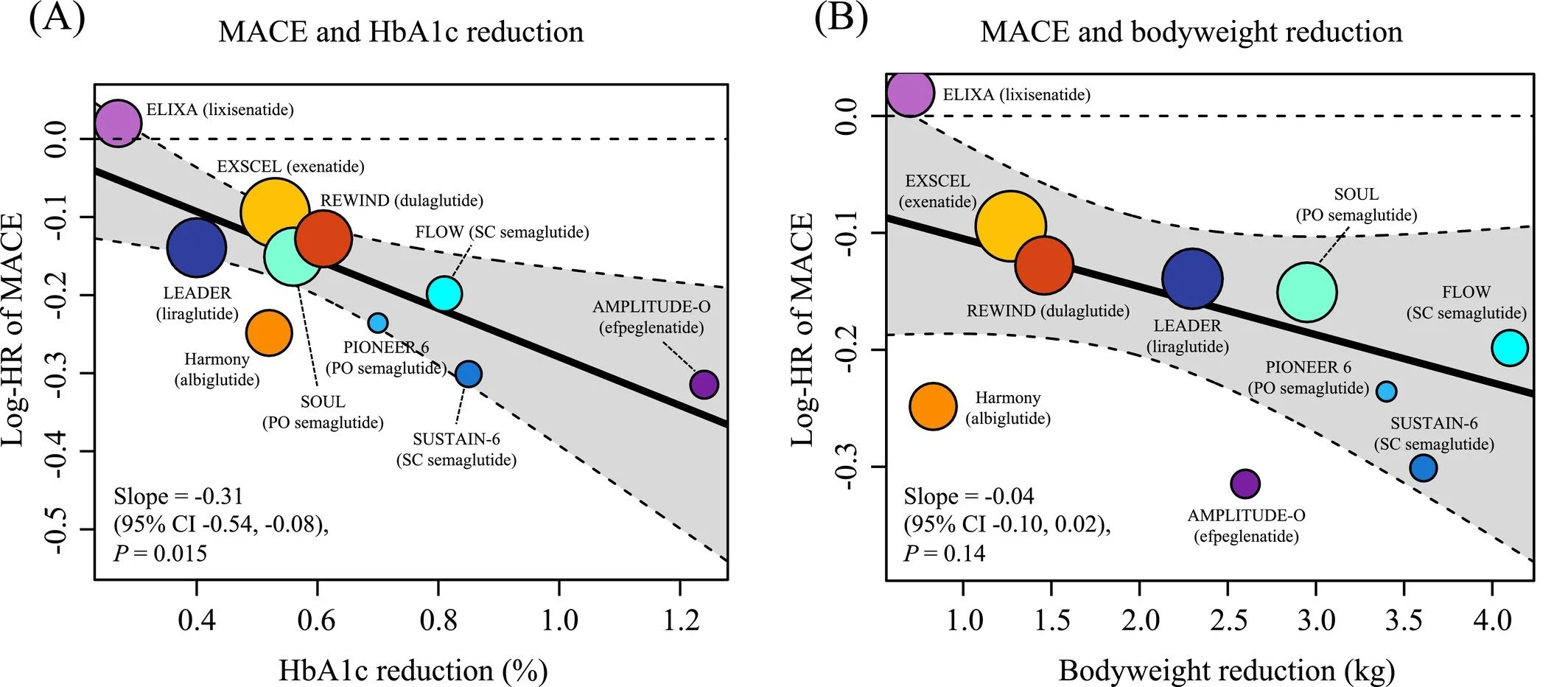
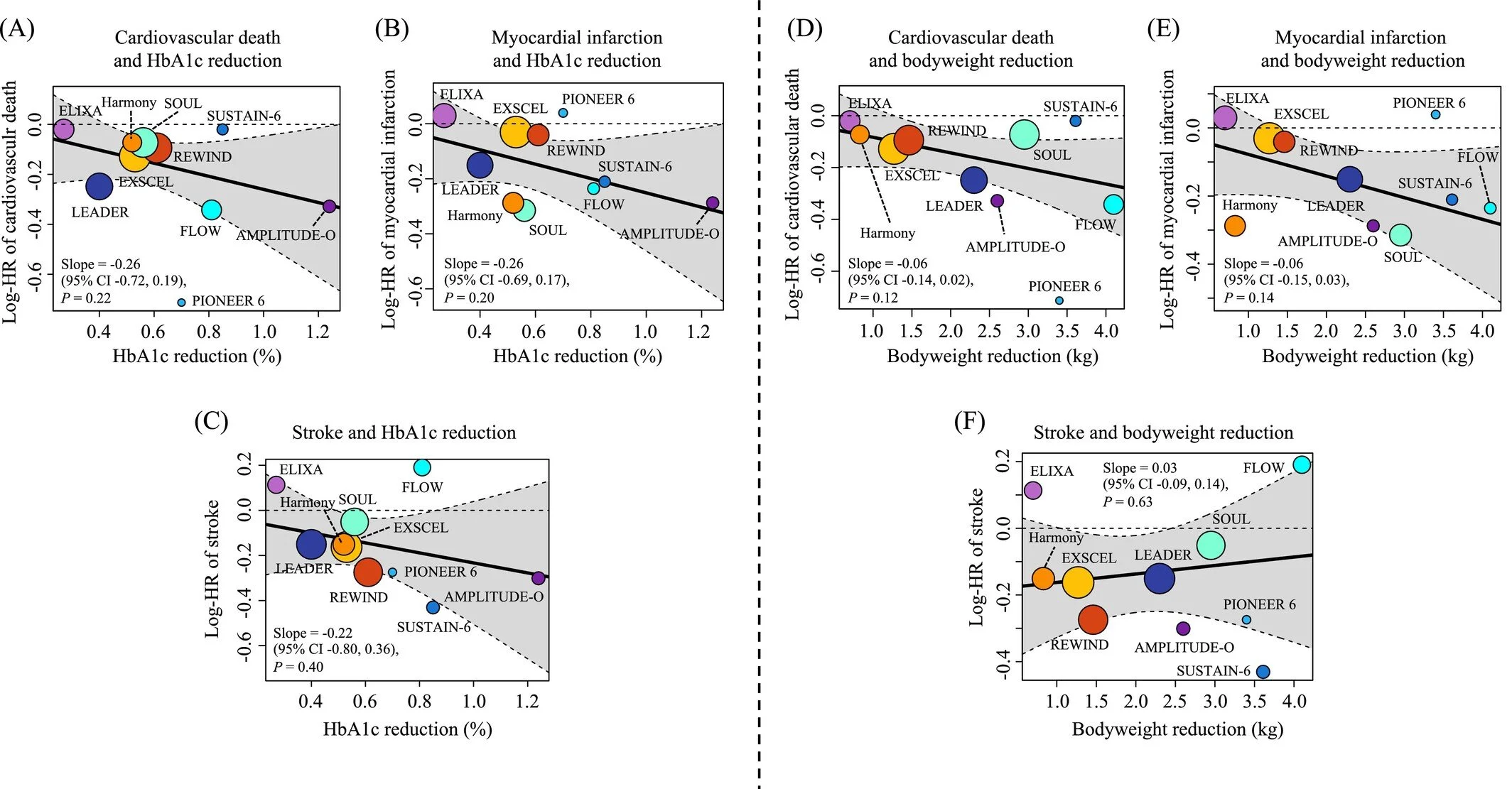
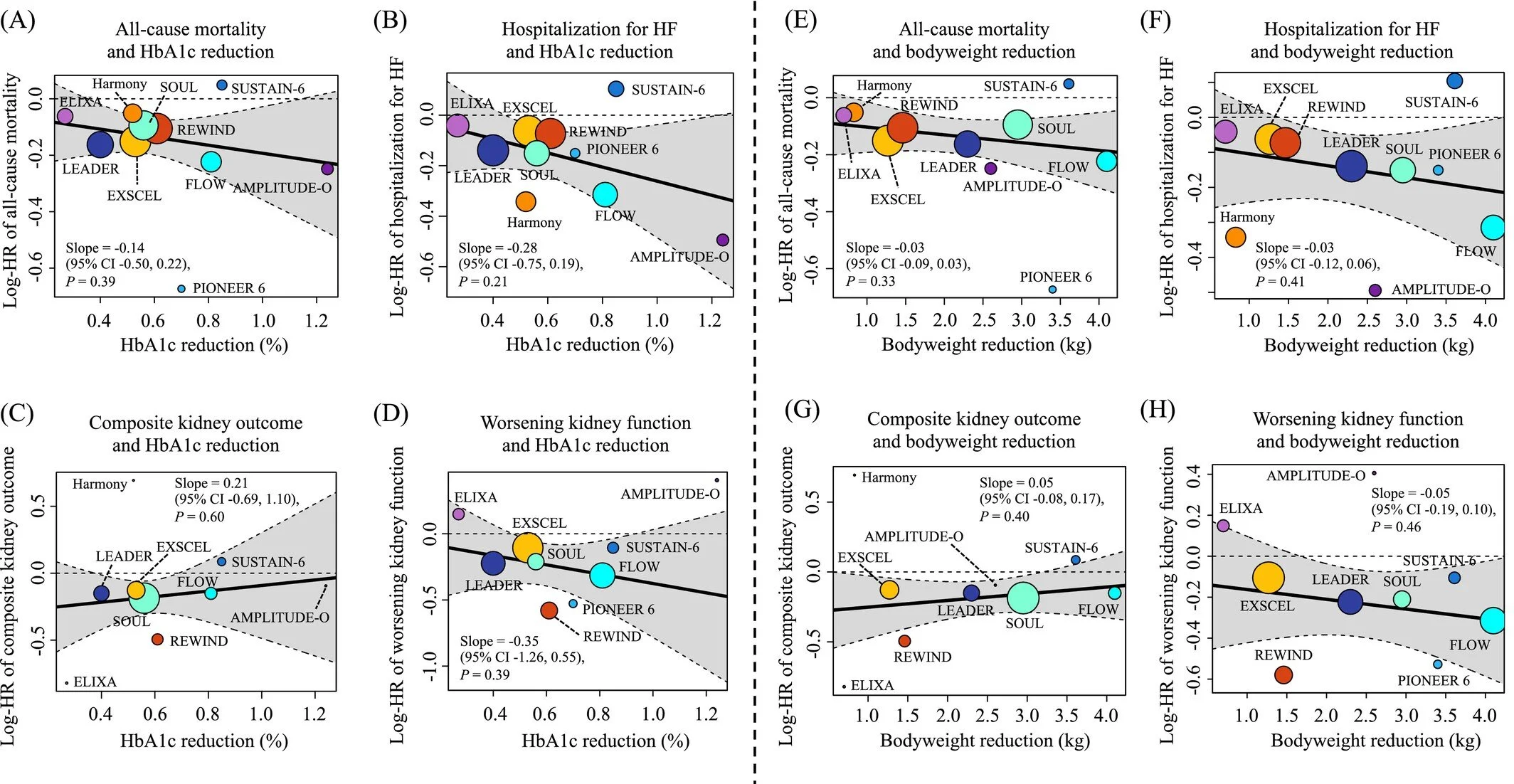
This updated meta‑regression analysis provides an in‑depth evaluation of how glucagon‑like peptide‑1 receptor agonists (GLP‑1RAs) reduce cardiovascular risk in individuals living with type 2 diabetes. Incorporating data from ten large randomized, placebo‑controlled trials—including the FLOW and SOUL trials—the study aggregates outcomes from 73,263 participants to assess how improvements in glycemic control relate to major cardiovascular benefits.
The authors report that GLP‑1RAs reduce major adverse cardiovascular events (MACE) by 14%, alongside significant reductions in hospitalization for heart failure and composite kidney outcomes. Notably, the analysis reveals that each 1% additional reduction in HbA1c corresponds to a 27% lower hazard ratio for MACE, underscoring the central role of glycemic improvement in cardiovascular protection.
Key Findings
GLP‑1RAs consistently reduce cardiovascular risk, demonstrating benefits beyond glucose lowering alone.
HbA1c reduction is strongly associated with improved cardiovascular outcomes, while weight reduction does not show an independent association in this analysis.
The inclusion of recent FLOW and SOUL trial data strengthens the robustness of the meta‑regression, providing updated evidence relevant to modern clinical practice.
CERC Contributions
We are proud to recognize the involvement of three CERC members in this publication:
Masashi Hasebe, co‑author and contributor to data synthesis and interpretation.
Satoshi Yoshiji, corresponding author, bringing expertise in endocrinology, genomics, and cardiometabolic disease within the CERC program.
Chen‑Yang Su, contributor to statistical analyses and quantitative methods through the Quantitative Life Sciences program.
Their collaboration reflects CERC’s commitment to advancing precision medicine and improving clinical outcomes for metabolic diseases.
CERC publication in Nature Metabolism


We’re delighted to spotlight a new publication in Nature Metabolism featuring contributions from our CERC community: Unravelling the molecular mechanisms causal to type 2 diabetes across global populations and disease‑relevant tissues. Congratulations to all collaborators—and in particular to CERC’s Satoshi Yoshiji and Chen‑Yang Su—for their role in this ambitious, international effort to decode the molecular mechanisms that drive type 2 diabetes (T2D).
What the study shows
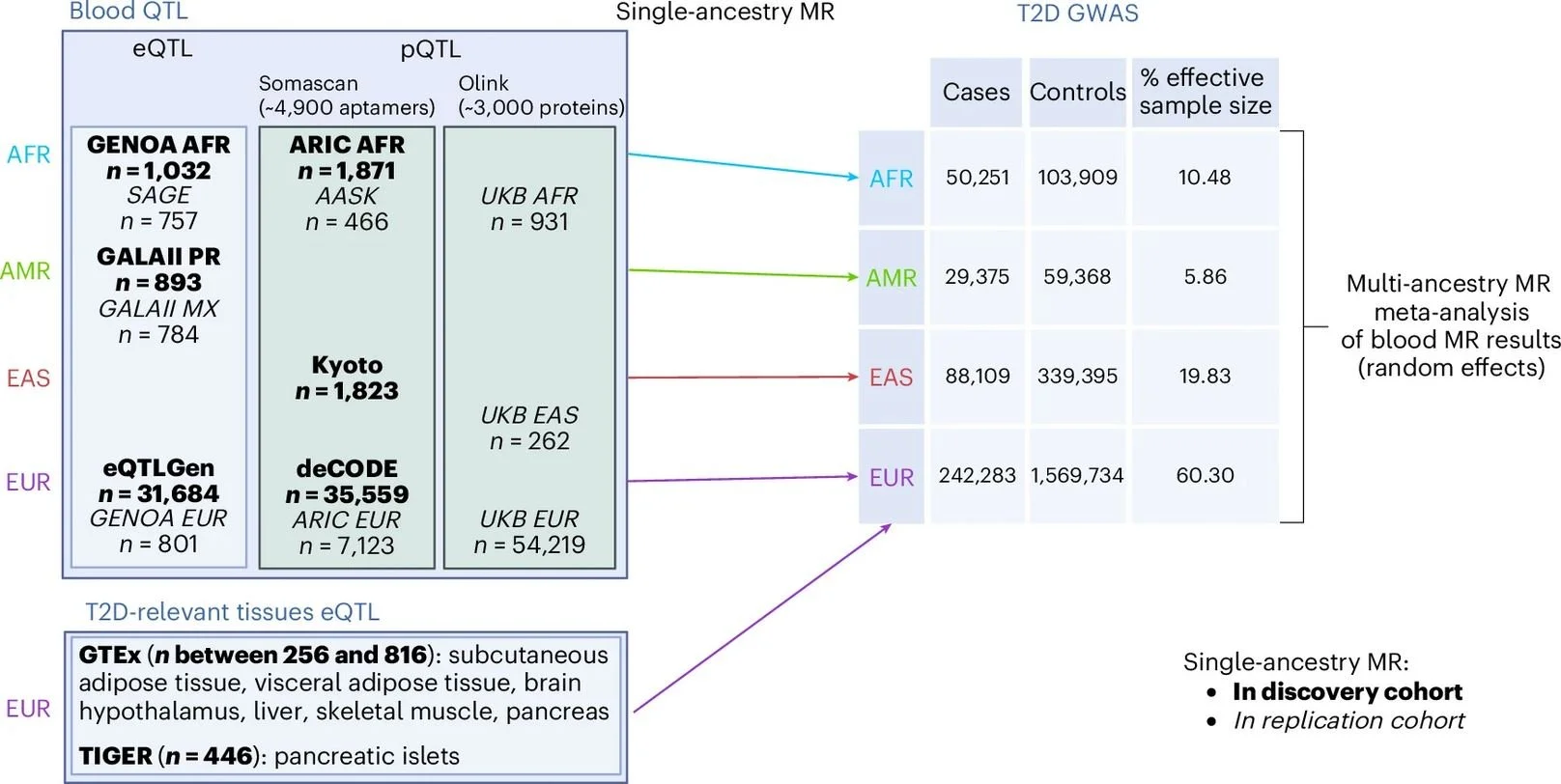
This work integrates large-scale genetics with multi‑omics across four global ancestry groups and seven T2D‑relevant tissues to pinpoint genes and proteins with causal effects on T2D risk. Using two‑sample Mendelian randomization and rigorous colocalization, the authors report causal links for 335 genes and 46 proteins based on blood molecular QTLs, and—crucially—identify 676 genes when expanding to tissues such as pancreatic islets, adipose (subcutaneous and visceral), liver, skeletal muscle, and hypothalamus.
By leveraging summary statistics from the Type 2 Diabetes Global Genomics Initiative—>2.5 million individuals, including >700,000 of non‑European ancestry—the team demonstrates that many causal signals are shared across ancestries yet highly heterogeneous across tissues. Analyses restricted to blood alone would have missed the majority of tissue‑specific signals, underscoring why disease‑relevant tissues matter for mechanism discovery and translational targeting.
Why it matters
Tissue context is critical. Only a small fraction of genes with causal effects in key T2D tissues show corresponding signals in blood, a finding that reframes how we prioritize therapeutic targets and biomarkers for metabolic disease.
Global representation strengthens discovery. Cross‑ancestry analyses both replicate shared mechanisms and reveal additional candidates observable only when genetic diversity is included—moving the field toward more equitable precision medicine.
CERC contributors
This publication includes contributions from Satoshi Yoshiji (Assistant Professor, Human Genetics; CERC in Genomic Medicine) and Chen‑Yang Su (former CERC doctoral student now postdoctoral researcher within the team). Their ongoing work in multi‑omics and genetic epidemiology continues to advance our program’s mission to translate human genetics into precision medicine for complex diseases.
Please join us in congratulating Satoshi, Chen‑Yang, and all collaborators on this milestone achievement!
Celebrating a New CERC Publication in Nature Communications









We are pleased to highlight a significant accomplishment within our research community: the publication of a new article in Nature Communications led by co‑first author Peyton McClelland. Congratulations to Peyton and to all collaborators involved in this impressive achievement!
Their study “A multi-ancestry genetic reference for the Quebec population”, published in Nature Communications on 31 January 2026, features contributions from several members of the CERC, including Alejandro Mejia Garcia, Mohadese Sayahian Dehkordi, Hongyu Xiao, Justin Pelletier, Vincent Chapdelaine, Geneviève Gagnon, Claude Bhérer and Daniel Taliun.
The article is available here: https://www.nature.com/articles/s41467-026-68820-7
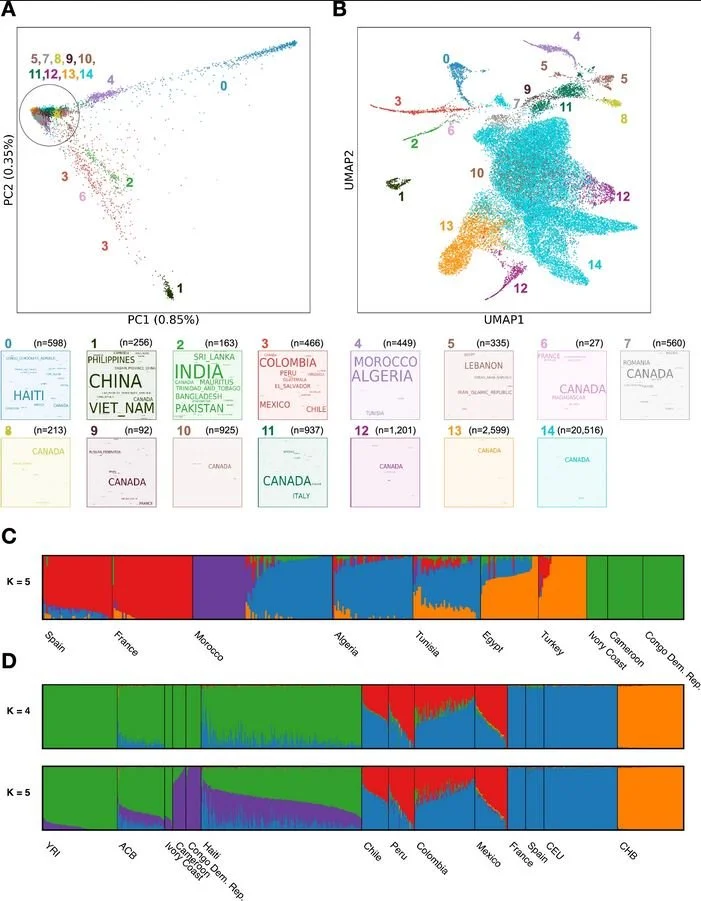
A multi-ancestry genetic reference for the Quebec population
The paper establishes a multi‑ancestry genetic reference for the Quebec population by analyzing genome‑wide genotypes from 29,337 CARTaGENE participants and whole‑genome sequencing of 2,173 individuals with grandparental origins in Canada, Haiti, and Morocco. This resource improves downstream analyses in several ways: the authors validate a Quebec‑specific imputation panel built from these sequences and show that, across 42 clinically relevant traits, it increases the number of associated loci by ~7% compared with using the larger TOPMed panel alone. The study also provides allele frequency data and GWAS results via public portals, enabling global reuse, and illustrates how the reference helps interpret variants in clinically important genes (e.g., SPG7). Together, the data and tools strengthen precision‑medicine efforts by better capturing Quebec’s demographic history and genetic diversity.
Please join us in congratulating Peyton and the entire team on this milestone achievement!
Congratulations on the PheWeb2 Publication in Nature Genetics




We are delighted to celebrate an important achievement within our research community: the publication of the PheWeb2 brief paper in Nature Genetics. Congratulations to Hongyu Xiao, Mehrdad Kazemi, Seyla Wickramasinghe and Daniel Taliun, as well as their collaborators, for this outstanding accomplishment!
Their article, “Exploring and visualizing stratified GWAS results with PheWeb2”, was published on 18 January 2026 in Nature Genetics (DOI: 10.1038/s41588-025-02469-8).
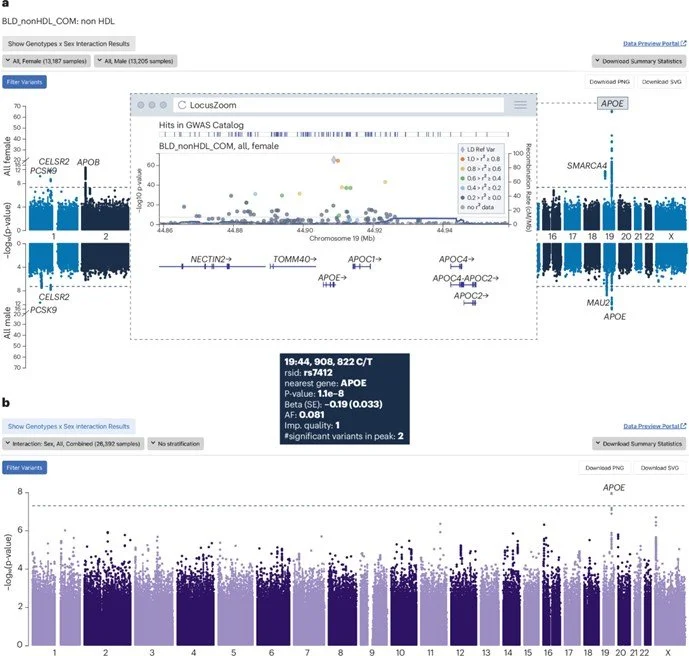
Advancing the Exploration of Stratified GWAS Data
PheWeb2 represents a complete redesign and enhancement of the original PheWeb tool, long used by the genetics community to navigate and share large-scale genome‑wide association study (GWAS) results. The new version addresses a major bottleneck in current research: the lack of intuitive tools for exploring sex‑stratified, ancestry‑stratified, and other stratified GWAS summary statistics.
The redesigned platform introduces major improvements, including:
Interactive visual comparisons such as Miami plots and stacked LocusZoom plots for side‑by‑side analysis of stratified GWAS or PheWAS results.
Support for numerous types of stratification, including genetic ancestry groups and sex‑specific analyses.
A modernized software architecture, decoupling the API from the user interface to improve maintainability and facilitate integration with external tools.
A fully updated UI built with Vue 3 and Vite, enhancing performance and ease of feature development.
These advances make PheWeb2 a powerful tool for researchers aiming to uncover subtle genetic effects that vary between population groups, ultimately helping drive new biological insights from increasingly diverse genomic datasets.
Please join us in congratulating Hongyu, Mehrdad, Seyla and Daniel on this exciting achievement!
Advancing Equity in Genetics and Genomics in Canada (AEG2C) Symposium Report Summary now available !
In November 2022, the Canadian Institutes of Health Research Institute of Genetics (CIHR-IG) and the McGill CERC in Genomic Medicine, in partnership with Genome Canada and Genome Québec, organized an international two-day online symposium on Equity, Diversity and Inclusion (EDI) in Genomics entitled: "Advancing Equity in Genetics and Genomics in Canada (AEG2C Symposium)".
The AEG2C symposium highlighted the importance and value of inclusion and diversity in genomics to advance our understanding of disease mechanisms and promote health equity. Members of the genetic and genomic community shared best practices for building a research enterprise that is diverse, inclusive, and beneficial to all, emphasizing that equitable approaches lead to better health outcomes. Lessons learned from international initiatives and Canadian studies were showcased, highlighting the need to engage diverse populations, seek input from multiple perspectives, and tailor policies and strategies to individual needs.
The full Report Summary is now available upon request at: info@cihr-irsc.gc.ca. More information here.
What's Next?
Building on the success and learnings inspired by this first edition, a second AEG2C symposium will be held on October 19-20, 2026. More information will be shared in the coming months. Stay tuned!
CERC student work highlighted by the American Journal of Human Genetics (AJHG)
Alejandro Mejia Garcia, a CERC PhD student under the supervision of Sirui Zhou was recently interviewed by The American Journal of Human Genetics (AJHG).
Each month, the editors of The American Journal of Human Genetics interview an author of a recently published paper. This month, they checked in with Alejandro to discuss his recent paper, “Using the ancestral recombination graph to study the history of rare variants in founder populations.”
You will be able to read the full interview here.
CERC student awarded the Spotlight Award - Confident Delivery at the Resonance event
On December 3rd, 2024 took place the inaugural Resonance Public Speaking Program, a new initiative designed by Drs. David Langlais and Andrew Churchill, together with a team of experiences trainees, to help students communicate their science clearly, confidently, and across disciplines.
Over ten weeks, participants worked with their coaches to refine their messages, sharpen delivery, and practise connecting with diverse scientific audiences across the VDIGM. The semester concluded with a 5-minute talk showcase that highlighted the breadth of research across our institute—from computational genomics to immunity and molecular engineering.
We would like to congratulate Thomas Zheng , a CERC PhD student under the supervision of Sirui Zhou for winning the Spotlight Award — Confident Delivery.
We also would like to say thanks to Tanja Sack, a CERC postdoctoral fellow in the Cuella Martin Lab for her coaching time.
3 CERC students received Excellence Awards from the Department of Human Genetics
We are proud to announce that 3 CERC students received Excellence Awards from the Department of Human Genetics. The Award Committee evaluated students’ contributions while enrolled in graduate studies in the Department of Human Genetics for 2025, including but not limited to academic performance and contributions to academic activities. Those students have made many important contributions locally, nationally and internationally in the form of talks, posters and publications. The 3 CERC nominees are:
Best MSc Publication Award
Cabré Romans, Júlia – Raquel Cuella Martin & Claude Bhérer Laboratories
Excellence Awards
Mejia Garcia, Alejandro (Ph.D. 3) – Sirui Zhou Laboratory
McClelland, Peyton (Ph.D. 4) – Daniel Taliun & Claude Bhérer Laboratories
Congratulations to the three of them !!!